Difference between revisions of "deLemus"
| Line 490: | Line 490: | ||
<htmltag tagname="img" src="https://wiki.laviebay.hkust.edu.hk/deLemus/RESEARCH_TEAMS/images/PublishedPlot/S2.png" alt="test for htmltag img" class="wikimg" style="display: block;width:70%;margin-left: auto;margin-right: auto;"></htmltag> | <htmltag tagname="img" src="https://wiki.laviebay.hkust.edu.hk/deLemus/RESEARCH_TEAMS/images/PublishedPlot/S2.png" alt="test for htmltag img" class="wikimg" style="display: block;width:70%;margin-left: auto;margin-right: auto;"></htmltag> | ||
| − | + | --> | |
== '''Deep Mutational Scanning Data''' == | == '''Deep Mutational Scanning Data''' == | ||
<big>The RBD-ACE2 binding data</big><ref>Greaney AJ, Starr TN, Gilchuk P, Zost SJ, Binshtein E, Loes AN, Hilton SK, Huddleston J, Eguia R, Crawford KHD, Dingens AS, Nargi RS, Sutton RE, Suryadevara N, Rothlauf PW, Liu Z, Whelan SPJ, Carnahan RH, Crowe JE Jr, Bloom JD. Complete Mapping of Mutations to the SARS-CoV-2 Spike Receptor-Binding Domain that Escape Antibody Recognition. Cell Host Microbe. 2021 Jan 13;29(1):44-57.e9. doi: 10.1016/j.chom.2020.11.007. Epub 2020 Nov 19. PMID: 33259788; PMCID: PMC7676316.</ref> <big>showed that R346S, N354S, E484R and S494P are the mutations lead to increased binding affinity in all the 5 background sequence.</big> | <big>The RBD-ACE2 binding data</big><ref>Greaney AJ, Starr TN, Gilchuk P, Zost SJ, Binshtein E, Loes AN, Hilton SK, Huddleston J, Eguia R, Crawford KHD, Dingens AS, Nargi RS, Sutton RE, Suryadevara N, Rothlauf PW, Liu Z, Whelan SPJ, Carnahan RH, Crowe JE Jr, Bloom JD. Complete Mapping of Mutations to the SARS-CoV-2 Spike Receptor-Binding Domain that Escape Antibody Recognition. Cell Host Microbe. 2021 Jan 13;29(1):44-57.e9. doi: 10.1016/j.chom.2020.11.007. Epub 2020 Nov 19. PMID: 33259788; PMCID: PMC7676316.</ref> <big>showed that R346S, N354S, E484R and S494P are the mutations lead to increased binding affinity in all the 5 background sequence.</big> | ||
| Line 581: | Line 581: | ||
|} | |} | ||
<big>Overall, by the first half of this year, '''R346S''' and '''E484R''' are the most potential dangerous mutations we captured.</big> | <big>Overall, by the first half of this year, '''R346S''' and '''E484R''' are the most potential dangerous mutations we captured.</big> | ||
| − | + | ||
==References== | ==References== | ||
<references> | <references> | ||
Revision as of 11:13, 9 June 2023
Dynamic Expedition of Leading Mutations in SARS-CoV-2 Spike Glycoproteins
The dynamic epidemiology of coronavirus disease 2019 (COVID-19) since its outbreak has been a result of the continuous evolution of its etiological agent, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Within the first 2 years of this pandemic, the World Health Organization (WHO) has already announced 4 variants of concern (VOC), namely alpha (B.1.1.7), beta (B.1.351), gamma (P.1), and delta (B.1.617.2), together with numerous variants of interest (VOI). The latest lineage to be designated a VOC would be omicron (B.1.1.529),[1] from which a diverse variant soup is generated.[2] From the original BA.1 strain of November 2021 to the most recent XBB and BQ.1 strains of late 2022,[3][4] each omicron subvariant has successively proliferated and outcompeted its once dominant antecedent.[5] The emergence of all these variants has brought along many novel mutations that continue to fine-tune the fitness of the virus,[6][7] leading to its persistent global circulation. Recent emerging variant (EV) data retrieved from GISAID, as of 17 January 2023, has revealed that the top 4 most rapidly spreading lineages are the BA.1.1.22, CH.1.1, XBB.1.5, and BQ.1.1 variants, among which XBB.1.5 has been found to be especially prevalent in the US,[8] making up of more than 40% of its sequence coverage in early January 2023.
Spike Glycoprotein
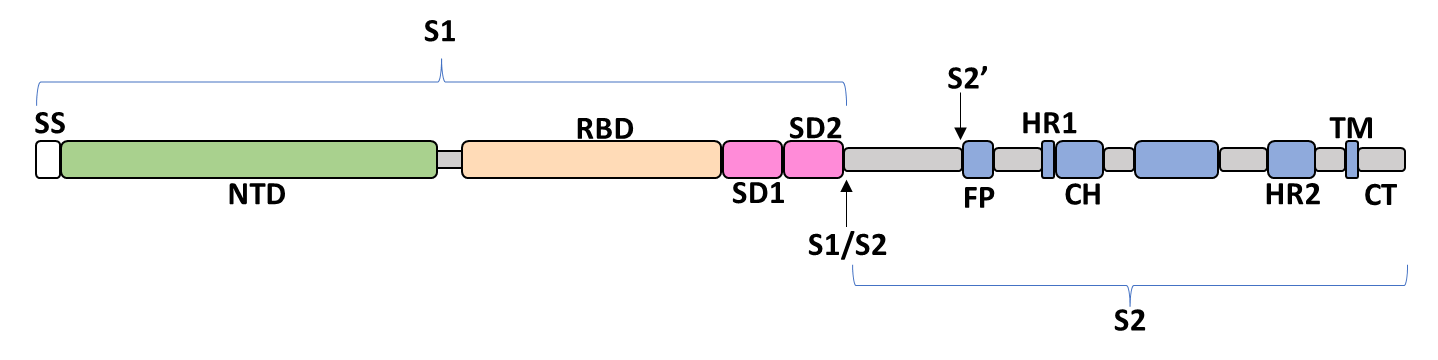
The spike glycoprotein of SARS-CoV-2 is a trimeric type I viral fusion protein that binds the virus to the angiotensin-converting enzyme 2 (ACE2) receptor of a host cell.[9] It is composed of 2 subunits: the N-terminal subunit 1 (S1) and C-terminal subunit 2 (S2), within which multiple domains lie. The S1 region facilitates ACE2 binding and is made up of an N-terminal domain (NTD), a receptor-binding domain (RBD), and 2 C-terminal subdomains (CTD1 and CTD2), while the downstream S2 region is responsible for mediating virus-host cell membrane fusion.
Update
The identified leading mutations in 2023 are listed as follows [10]:

- Generated 3D structure of spike protein with highlighted leading mutations (AlphaFold2, colab version 2022).
Here are the recently confirmed leading mutations.
2023.05.01 - 2023.05.12
| Outlined Mutations | Confirmed in VOC/Emerging Variants |
|---|---|
| F490P | XBB.1.9.1 |
| L858I | CH.1.1.1 |
*The reported mutations of detected variants are from GISAID[11]

- Generated 3D structure of spike protein with highlighted leading mutations (AlphaFold2, colab version 2022).
Here are the recently confirmed leading mutations.
2023.05.01 - 2023.05.12
| Outlined Mutations | Confirmed in VOC/Emerging Variants |
|---|---|
| F456L | FD.1.1 |
| S494P | XBB.2.3 & XBB.1.1 |
| T572I | FY.1 ( XBB.1.22.1.1 ) |
*The reported mutations of detected variants are from GISAID

- Generated 3D structure of spike protein with highlighted leading mutations (AlphaFold2, colab version 2022).
Here are the recently confirmed leading mutations.
2023.04.01 - 2023.04.21
| Outlined Mutations | Confirmed in VOC/Emerging Variants |
|---|---|
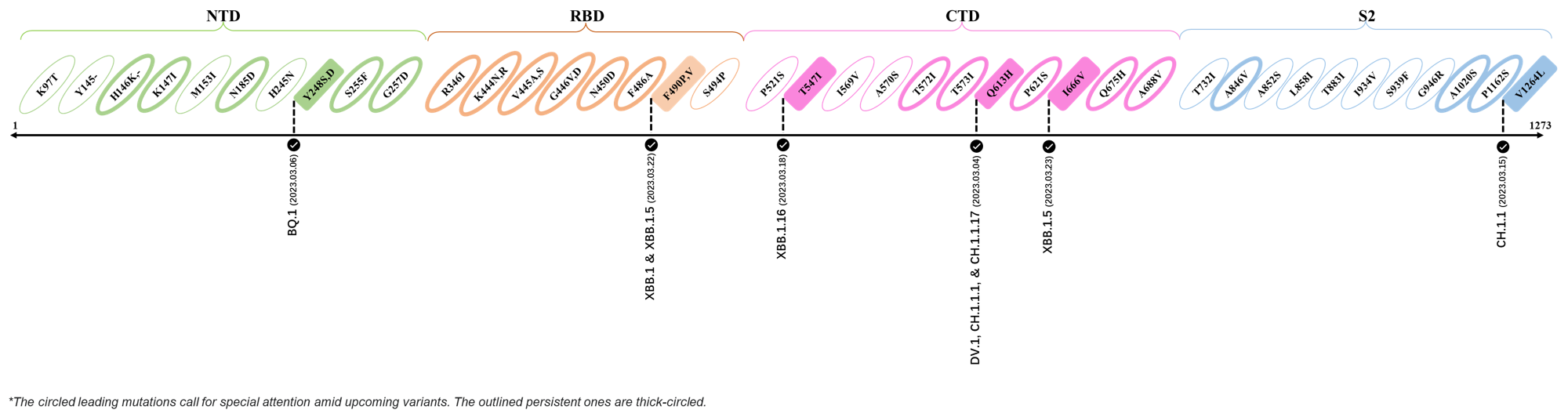
| H146K | XBB.1.5 & XBB.1.16 |
| M153I | XBB.2.3.3 |
| E180V | XBB.1.16 |
| K444R | XBB.1.5 |
| T478R | XBB.1.16, XBB.1.5, CH.1.1.2 & XBB.2.3 |
| F490P | XBB.2.6 |
| S494P | XBB.1.5 |
| Q613H | XBB.1.16 |
| P621S | XBB.2.3 |
| A688V | XAY.1.1.1 |
*The reported mutations of detected variants are from Cov-Lineages[12]
- Generated 3D structure of spike protein with highlighted leading mutations (AlphaFold2, colab version 2022).
Here are the recently confirmed leading mutations.
2023.03.01 - 2023.03.21
| Outlined Mutations | Confirmed in VOC/Emerging Variants |
|---|---|
| Y248S | BQ.1 |
| F490P | XBB.1 & XBB.1.5 |
| T547I | XBB.1.16 |
| Q613H | DV.1, CH.1.1.1 & CH.1.1.17 |
| I666V | XBB.1.5 |
| V1264L | CH.1.1 |
- Generated 3D structure of spike protein with highlighted leading mutations (AlphaFold2, colab version 2022).
Here are the recently confirmed leading mutations.
2023.02.03 - 2023.02.20
| Outlined Mutations | Confirmed in VOC/Emerging Variants |
|---|---|
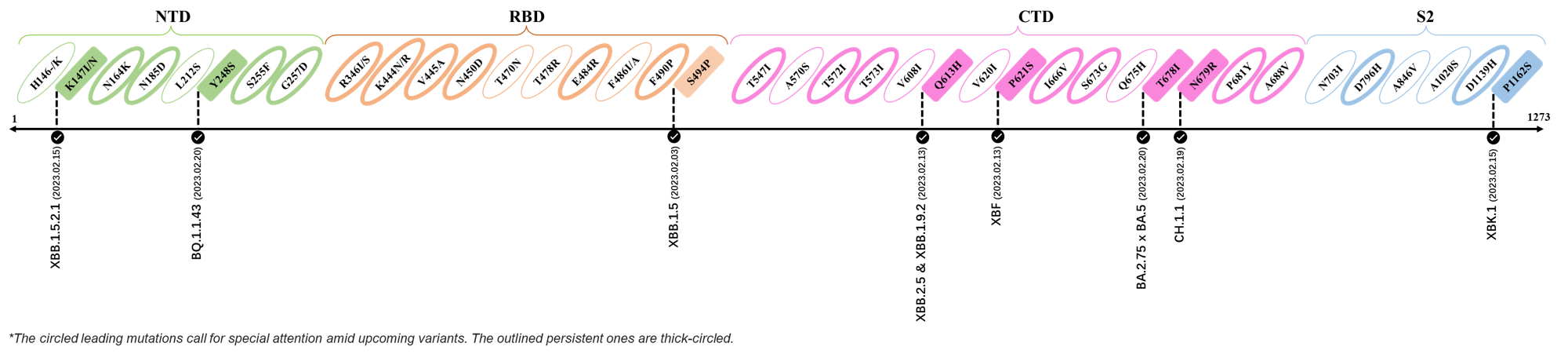
| K147I | XBB.1.5.2.1 |
| Y248S | BQ.1.1.43 |
| S494P | XBB.1.5 |
| Q613H | XBB.1.9.2 & XBB.2.4 |
| P612S | XBF |
| T678I | BA.2.75 x BA.5 |
| N679R | CH.1.1 |
| P1162S | XBK.1 |
- Generated 3D structure of spike protein with highlighted leading mutations (AlphaFold2, colab version 2022).
Here are the recently confirmed leading mutations.
2023.01.31
| Outlined Mutations | Confirmed in VOC/Emerging Variants |
|---|---|
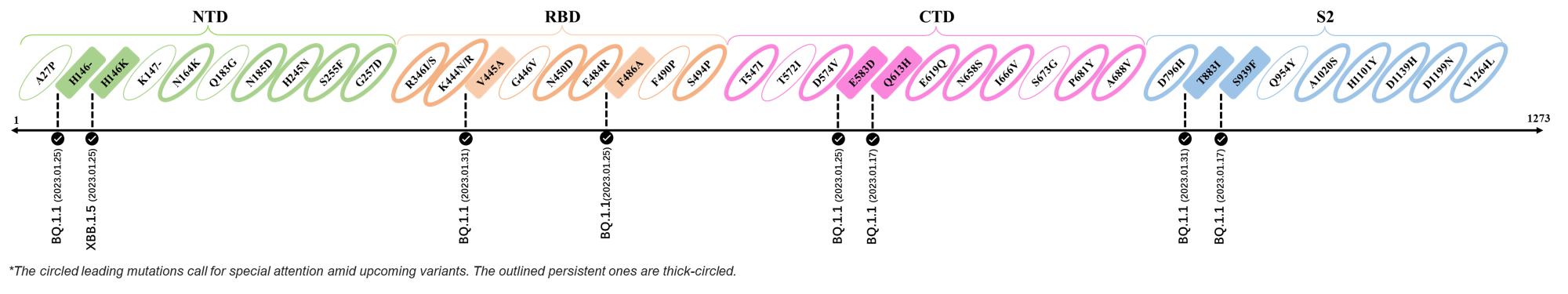
| V445A | BQ.1.1 |
| T883I | BQ.1.1 |
2023.01.17 - 2023.01.25
| Outlined Mutations | Confirmed in VOC/Emerging Variants |
|---|---|
| H146- / H146K | BQ.1.1 / XBB.1.5 |
| F486A | BQ.1.1 |
| E583D | BQ.1.1 |
| Q613H | BQ.1.1 |
| S939F | BQ.1.1 |
Deep Mutational Scanning Data
The RBD-ACE2 binding data[13] showed that R346S, N354S, E484R and S494P are the mutations lead to increased binding affinity in all the 5 background sequence.
| Unique Mutations | Date | Wuhan | Alpha | Beta | Eta | Delta |
| R346S | 2023.01 | 0.12 | 0.14 | 0.07 | 0.03 | 0.11 |
| N354S | 2023.05 | 0.03 | 0.01 | 0.04 | 0.32 | 0.02 |
| E484R | 2023.01 | 0.06 | 0.04 | - | - | 0.11 |
| S494P | 2023.01 | 0.33 | 0.18 | 0.13 | 0.14 | 0.06 |
Immune escape data[14] shows that the escape ability of R346S, V445A, G446I, and E484R against certain antibodies exceeds 90% mutations.
| Unique Mutations | Date | Antybody1 | Antybody2 | Antybody3 | Antybody4 | Antybody5 |
| R346S | 2023.01 | COV2-2082 | COV2-2096 | COV2-2479 | COV2-2832 | |
| V445A | 2023.01 | COV2-2050 | COV2-2094 | COV2-2479 | COV2-2499 | COV2-2677 |
| G446I | 2023.05 | COV2-2096 | COV2-2479 | COV2-2499 | ||
| E484R | 2023.01 | COV2-2050 | COV2-2096 | COV2-2479 | COV2-2832 |
Overall, by the first half of this year, R346S and E484R are the most potential dangerous mutations we captured.
References
- ↑ Karim, S. S. A. & Karim, Q. A. Omicron SARS-CoV-2 variant: A new chapter in the COVID-19 pandemic. Lancet 398, 2126 (2021).
- ↑ Callaway, E. COVID ‘variant soup’ is making winter surges hard to predict. Nature 611, 213 (2022).
- ↑ Wang, Q. et al. Alarming antibody evasion properties of rising SARS-CoV-2 BQ and XBB subvariants. Cell 186, 279 (2023).
- ↑ Qu, P. et al. Enhanced Neutralization Resistance of SARS-CoV-2 Omicron Subvariants BQ.1, BQ.1.1, BA.4.6, BF.7, and BA.2.75.2. Cell Host Microbe 31, 9 (2023)
- ↑ Rössler, A. et al. BA.2 and BA.5 Omicron Differ Immunologically from Both BA.1 Omicron and Pre-Omicron Variants. Nat Commun 13, 7701 (2022)
- ↑ Carabelli, A. M. et al. SARS-CoV-2 variant biology: Immune escape, transmission and fitness. Nat Rev Microbiol (2023). DOI: https://doi.org/10.1038/s41579-022-00841-7.
- ↑ Witte, L. et al. Epistasis lowers the genetic barrier to SARS-CoV-2 neutralizing antibody escape. Nat Commun 14, 302 (2023).
- ↑ Callaway, E. Coronavirus variant XBB.1.5 rises in the United States — is it a global threat? Nature 613, 222 (2023).
- ↑ Jackson, C. B., Farzan, M., Chen, B. & Choe, H. Mechanisms of SARS-CoV-2 entry into cells. Nat Rev Mol Cell Biol 23, 3 (2021).
- ↑ deLemus team, Analysis of Leading Mutations in SARS-CoV-2 Spike Glycoproteins (in preparation, 2023).
- ↑ GISAID https://gisaid.org/
- ↑ Cov-Lineages https://cov-lineages.org/
- ↑ Greaney AJ, Starr TN, Gilchuk P, Zost SJ, Binshtein E, Loes AN, Hilton SK, Huddleston J, Eguia R, Crawford KHD, Dingens AS, Nargi RS, Sutton RE, Suryadevara N, Rothlauf PW, Liu Z, Whelan SPJ, Carnahan RH, Crowe JE Jr, Bloom JD. Complete Mapping of Mutations to the SARS-CoV-2 Spike Receptor-Binding Domain that Escape Antibody Recognition. Cell Host Microbe. 2021 Jan 13;29(1):44-57.e9. doi: 10.1016/j.chom.2020.11.007. Epub 2020 Nov 19. PMID: 33259788; PMCID: PMC7676316.
- ↑ Tyler N. Starr., et al., Shifting mutational constraints in the SARS-CoV-2 receptor-binding domain during viral evolution.Science377,420-424(2022).DOI:10.1126/science.abo7896
